Competitiveness of Our Technology
01
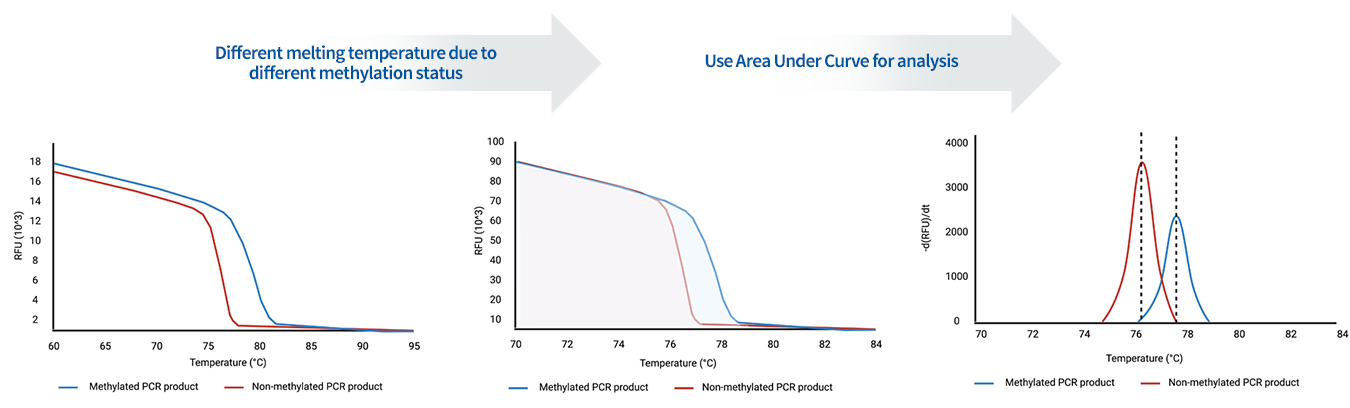
LepiMA is an upgrade of the conventional
MS-HRM and amplifies more of the methylated
DNA to allow more sensitive
and specific detection of methylated DNA.
MS-HRM and amplifies more of the methylated
DNA to allow more sensitive
and specific detection of methylated DNA.
02
LepiMA can be applied to diverse
methylation markers and allows calculation
of the ratio between methylated
and non-methylated DNA.
methylation markers and allows calculation
of the ratio between methylated
and non-methylated DNA.